Searching Genbank
Go to parent Basic bioinformatics concepts, databases and tools#Exercises_during_the_training
NCBI maintains a large set of biological databases, including Genbank. Genbank contains annotated (sequences + info) primary (not curated) sequence data. The sequence records are structured in Genbank format.
![]() Warning: Information is liquid. Records change all the time: info is removed and added. Therefore screenshots may not be up-to-date.
Warning: Information is liquid. Records change all the time: info is removed and added. Therefore screenshots may not be up-to-date.
Contents
- 1 Searching Genbank
- 1.1 Exercise 1: human BRCA2
- 1.2 Exercise 2: presenilin 1
- 1.3 Exercise 3: 10 largest human mRNAs
- 1.4 Exercise 4: obtaining the most recent submissions
- 1.5 Exercise 5: Using Ugene to download records from Genbank
- 1.6 Exercise 6: Genbank searches on CLC Bio Main workbench (VIB only)
- 1.7 Exercise 7: genomic scaffold for Drosophila
Searching Genbank
Go to http://www.ncbi.nlm.nih.gov/.
Exercise 1: human BRCA2
Search for the human BRCA2 mRNA sequence.
Select Nucleotide in the database select box, type human BRCA2 in the search text box and click Search
The search results page shows by default chunks of 20 records, that fullfil your search criteria, with a summary of each record. This search generates more than 1000 records that all contain the words Homo sapiens and BRCA2. But not all of them contain the BRCA2 sequence.
For instance, the 18th and 19th contain the sequence of chromosomes of red deer.
Go to page 6 of the results:
- Type 6 in the Page box
- Click ENTER on your keyboard

Go to hit 102. Click the link:Mus musculus BRCA2 (Brca2) mRNA, complete cds to open
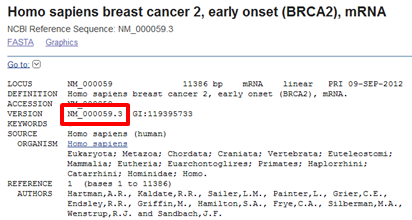
the corresponding Genbank record. The record is formatted by default in GenBank format. Genbank format divides the record into sections or fields with the sequence at the bottom of the record. Keywords like LOCUS, DEFINITION, ACCESSION... mark the content of the different sections.The sequence is indeed a BRCA2 sequence, however not from human but from mouse. When you look for Homo sapiens in this record using the Find function in your browser, you see that the record contains the expression Homo sapiens in the FEATURES section. In the note of the CDS feature it is stated that this gene is similar to Homo sapiens breast cancer susceptibility gene BRCA2. Because the record fullfils all the search criteria that were specified it is returned by the search:
- it contains the word BRCA2
- it contains the expression Homo sapiens
Go to page 14 of the results, to number 276 and click the link:
Homo sapiens zygote arrest 1-like (ZAR1L), RefSeqGene on chromosome 13
Look for the word BRCA2 in this record. In the FEATURES section of the record (in green) you see that the record contains only part of the BRCA2 sequence ! That's normal since this record is meant to hold the sequence of gene ZAR1L, the gene neighbouring BRCA2 on chromosome 13. But the record is returned by your search since it mentions the word BRCA2 (and Homo sapiens).
Feature annotation
The ZAR1L record contains a genomic sequence (a DNA sequence instead of a mRNA sequence). You can check the sequence type in the LOCUS section at the top of the record (in red).
Since genomic sequences contain introns, DNA sequences have more annotations than mRNA sequences. Let's zoom in on the gene annotation (green). NCBI's definition of gene is not very instructieve: region of biological interest identified as a gene and for which a name has been assigned. In Genbank
- gene features contain introns, exons and UTRs (untranslated regions)
- mRNA features contain exons and UTRs
- CDS (coding sequence) features contain exons only (so they start at ATG and end at the stop codon)
You can find a complete description of Genbank features here.
The text is green should be read as: This feature is a gene called BRCA2 located on the reverse strand from position 1475. The sequence of this gene is not fully contained in the sequence displayed in this record. In these annotations the following terminology is used:
- complement
The word complement e.g. in the gene annotation in green means that this feature, a gene called BRCA2, is located on the reverse strand (relative to the sequence shown at the bottom of the record). Records of mRNA sequences always contain the strand that contains the CDS. Records of DNA sequences contain the strand that was designated as the "+" strand so genes can be located on the strand that is shown but they can also be located on the reverse strand. - < or >
The BRCA2 gene starts at position 1475 and ends upstream of the sequence that is shown at the bottom of the record: this is what the minus sign in <1..1475 means. A < or > sign means that the record only contains a partial sequence. So the sequence at the bottom of the record contains only a part of the complementary strand of the BRCA2 gene. - join
Unlike mRNA records, which do not contain introns since they are spliced out from the mRNA, DNA records do contain intron sequences. Look at the annotation of the BRCA2 mRNA (in blue). It does contain introns which is indicated by join in complement(join(<428..533,1288..1475)). So the first exon is located from position 1475 to position 1288 on the reverse strand and the first intron is located from position 1287 to position 534.
Translation of your search into a query to the database
You can learn a lot from the translation of the search terms that you typed into the search box into the query that was effectively used to search the database (see in the right column of the results page under Search details).
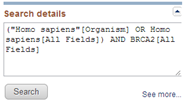
You see that indeed both the term BRCA2 and the expression Homo sapiens was searched in all fields of the records.
Similarly, when you go to the summary of hit 306, you see that the record contains the sequence of an interaction partner of BRCA2. So this record does not at all contain the BRCA2 sequence.
Limit the number of search results
Our search generates a lot of rubbish results, how can we avoid this ?
Advanced searches
A good way to limit the number of search results, is to use an advanced search. The Advanced link is located below the search term box

This leads you to the Nucleotide Advanced Search Builder and helps you to create complex queries.
- Select Organism in the first field box and type Homo sapiens in the corresponding search text box
- Select AND as boolean operator
- Select Gene Name in the second field box and type BRCA2 in the corresponding search text box
- Click Search
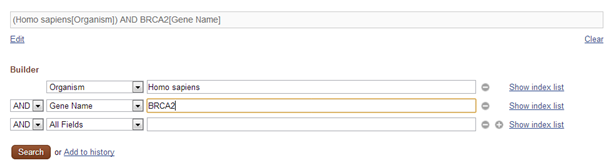
Now you are looking for records that contain human sequences (with the expression Homo sapiens in the Organism field specified by setting Organism to Homo sapiens) and that mention the word BRCA2 in the Gene Name field(specified by setting Gene Name to BRCA2). This limits the number of results significantly.
Filters
At the left side of the results summary page there's a list of filters you can use to restrict the search results even more. Filters allow you to restrict the search by date, organism, quality (RefSeq), sequence type and other characteristics. The number after the filter shows how many records remain after activating the filter.
At the right side you have an additional filter: Results by taxon. Remarkably not all records are human although you did specifically ask for that in the Advanced search. There is one record of a synthetic construct.

Click the synthetic construct filter to open the record.
| Look at the Organism field to see if it contains Homo sapiens |
|---|
 You don't see the expression Homo sapiens here. |
Then why was the record returned by the search ? Scroll down to the FEATURES section.
| Look at the source annotation to see if you find Homo sapiens |
|---|
 You see that there are two organisms annotated:
|
So searching the Organism field will search the Organism field in the general annotation but also the /organism fields in the feature annotation.
Activate the Homo sapiens filter in the Results by taxon section.
| How should you do the search to return human sequences without synthetic constructs? |
|---|
| Look at the translation of the query (with Homo sapiens filter):
 If you want to be sure not to return sequences from mixed origin (with multiple organisms) you have to search the Primary organism field: [porgn]. |
| Activate the mRNA filter to only see records that contain mRNA sequences |
|---|
| Click mRNA (red) in the filter section at the left.
 |
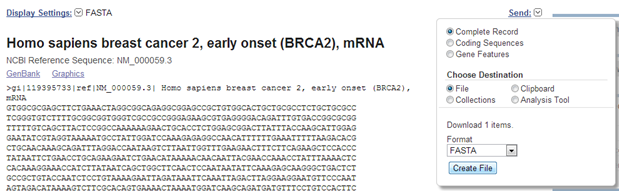
Click the first link:
Homo sapiens breast cancer 2, early onset (BRCA2), mRNA. This is the longest sequence. You can see this because the results summary page shows the lengths of the sequences under the links to the records.

RefSeq records
Note that this record comes from the RefSeq database. RefSeq is the non-redundant, curated subset of Genbank.
- Non-redundant means that there are no duplicates, each sequence is represented by a single record
- Curated means that the content is revised, most RefSeq records are created by NCBI staff based on the corresponding Genbank records to obtain the longest sequence with a minimum of sequencing errors and the most accurate annotation.
| Was this RefSeq record curated ? |
|---|
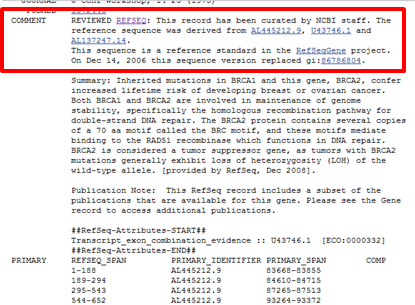
| You can verify this by checking the quality label in the COMMENT section of the Refseq record. You see that the label is REVIEWED REFSEQ: This record has been curated by NCBI staff. |
| What are the accession numbers of the original sequences from which this RefSeq record was derived ? |
|---|
| You can find this information in the COMMENTS section of the record, along with a link to the previous version of this record. |
| What are the synonyms of BRCA2 ? |
|---|
| You can find this information in the FEATURES section of the record. Go to the gene annotation. You can see in the /gene_synonym subsection that the synonyms of BRCA2 are BRCC2; BROVCA2; FACD; FAD; FAD1; FANCB; FANCD; FANCD1; GLM3; PNCA2 |
Note that you can select to download the coding sequence (CDS) alone.
| What is the location of the CDS? |
|---|
| In the FEATURES section you can find the CDS annotation. The numbers you see here: 228..10484 indicate the start and stop position of the coding part in the sequence that is displayed at the bottom of the record. So the sequence in this record contains more than only the coding sequence. This was to be expected since it is a mRNA sequence, which normally also contains the 5' and 3' UTR. |
Now search the RefSeq record of the non-predicted BRCA2 mRNA of dog (canis lupus).
| Is this RefSeq record curated by NCBI staff ? |
|---|
Do an Advanced search:
Activate the following filters:
You get three records, the title of two of them starts with the word "PREDICTED" so these will not be curated. In the COMMENTS section of the third record you see: The RefSeq annotation approach uses both collaborator supplied sequence information and automated BLAST analysis to provide an initial RefSeq record. Records are subject to validation to correct annotation errors and provide annotation in a more consistent format. Descriptive information, including Official Nomenclature and additional citations, are applied to the records. These initial records have a PROVISIONAL, PREDICTED, or INFERRED status. Additional manual curation is applied to this set of RefSeq records to provide the optimal sequence record, and to fix sequence errors including mis-association with a locus (as might occur for closely related gene families), chimeric sequences, vector or linker contamination, or apparent sequencing errors. Both the nucleotide and protein sequence record may change due to this process. Sequence level review is carried out primarily by NCBI staff but some records are provided via collaboration. These records have a VALIDATED status. Additional annotation, a summary description, and other functional information may be applied, as available, during the sequence review process. These records have a REVIEWED status." |
Exercise 2: presenilin 1
Search for presenilin 1 in the Nucleotide database. Make sure that you remove the filters from the previous exercise !
On the results page you see that the search generates more than 2000 hits, many of them being full chromosome or scaffold sequences.
Sequencing and assembly
Chromosome sequences are not sequenced as such due to size limitations of the sequencing process. They are chopped up into small overlapping fragments that are individually sequenced. This gives rise to a large set of short sequence reads (sequences of these small fragments) that are assembled into larger sequences. Assembly means that you look for overlaps between reads and use these overlaps to extend the sequence.
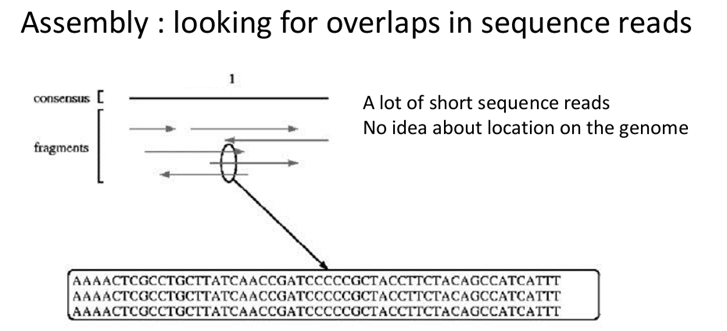
So chromosomes are fragmented into small overlapping fragments and these fragments are sequenced. Then the resulting reads are assembled to form chromosome sequences again.
In this way short reads are assembled into larger contigs, contigs are assembled into scaffolds and scaffolds are assembled into full chromosomes:
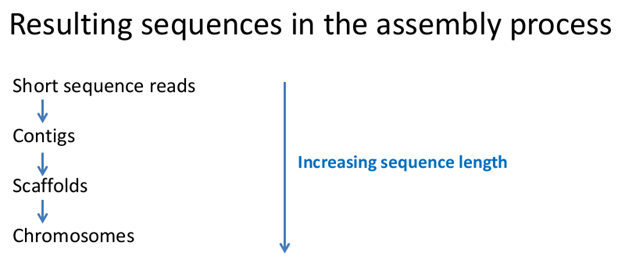
Although short sequences are to be removed from Genbank once longer assembled ones are available, the removal process takes time. Often both longer and smaller sequences exist in Genbank. So you can find a gene in many formats in Genbank: as a gene, a mRNA, a protein, a part of a scaffold, a part of a chromosome...
Limit the number of search results
We want to get rid of these long sequences using a more targeted search.We cannot search presenilin 1 in the gene name field as we have done for BRCA2 in the previous exercise. If you specify Gene Name as the field to search in an advanced search, the /gene= sections of the FEATURES field will be searched (see green and blue in the figure below):

According to the detailed description of Genbank features /gene= represents the symbol of the gene corresponding to a sequence region. Gene symbols are not full names (like presenilin 1) but abbreviatons consisting of three or four letters and an arabic number, like BRCA2.
We have to find another field to search in the list of available search fields. The most relevant fields to search are Protein Name and Title.
Now which do you choose?
Protein Name will limit the search to records that contain the expression presenilin 1 in the protein features annotations. Most scaffolds and chromosomes are annotated so they do contain protein feature annotations. Using this field will not remove scaffolds and chromosomes from the results.
Title will limit the search to records that contain presenilin 1 in the title, i.e. the description line that you can click on the results summary page and that appears at the top of the record. So using Title will get rid of all chromosome and scaffold sequences since they do not contain the names of genes or proteins in their description lines.
| Use an advanced search to search Genbank records with presenelin 1 in the Title field. |
|---|
| Do an Advanced search: Title: presenilin 1. |
This search still returns a lot of results.
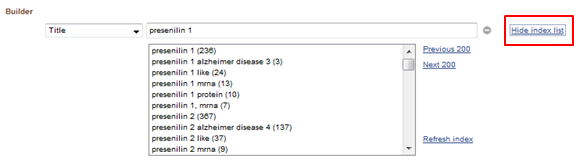
![]() If you're not sure how to enter a term (with/without dashes, with 1 attached to the name or not...), use the Index list to select the correct term
If you're not sure how to enter a term (with/without dashes, with 1 attached to the name or not...), use the Index list to select the correct term
| Restrict the search to mouse presenilin 1. |
|---|
Do an Advanced search:
|
This limits the number of results.
You do not need to use the Latin organism name. The search tool can translate English to Latin organism names (at least for the most used model organisms).
| Check the organism field of the two last hits containing the promoter sequences. |
|---|
| Organism = Mus sp. Click the Taxonomy link in the right menu to go the Taxonomy browser. |
| What does Mus sp. mean ? |
|---|
| The Comments and References section of the Taxonomy browser states that "Many of the sequence records listed under the "Mus sp." name are scanned in from journal articles that have identified the organism only with the english vernacular "mouse" or "mice". The source for the majority of these records is likely to be the house mouse "Mus musculus"." |
So these sequences are probably coming from the Mus musculus but you're not entirely sure.
| Repeat the search using Mus musculus. |
|---|
Do an Advanced search:
|
As expected, the results summary shows that the two Mus sp. records are not returned when you search for Mus musculus.
| From these results, select the RefSeq records. |
|---|
| Use the RefSeq filter in the Filter menu at the left of the page or look for accession numbers that contain an underscore (see slides). |
Exercise 3: 10 largest human mRNAs
| Do an advanced search to find the accession numbers of the 10 largest human mRNA sequences in Genbank ? |
|---|
|
The 10 largest human mRNAs all encode isoforms of titin, a giant protein that provides sctructure and elasticity of muscle cells.
Exercise 4: obtaining the most recent submissions
Many scientists regularly visit the NCBI website to check the most recent submissions on their favourite organism. In this case it's very handy if you can filter the recent submissions from the search results.
We will search for recent submissions on sugar cane (Saccharum officinarum) in the Nucleotide database.
| How many submissions were done of sugar cane sequences in 2017 ? |
|---|
You need an advanced search for this:
|
Exercise 5: Using Ugene to download records from Genbank
Ugene is free software to perform various bioinformatics analyses. It has similar functionality as the CLC Bio Main Workbench but it is free to use. It allows the retrieval of data from online databases, like Genbank. It provides tools for visualization of sequences, multiple sequence alignments, phylogenetic trees and 3D structures.
You can search the following fields in Genbank records:
- accession or version numbers
- author, gene and/or organism names
Retrieve sequences from Genbank based on accession numbers
We will use Ugene to retrieve record AC009453 from the human genome project.
Ugene has been installed on the BITS laptops. To open the software double click the ![]() icon on the desktop.
icon on the desktop.
To download a sequence from Genbank:
- Click File in the top menu
- Click Access remote database

Scroll down the Database dropdown menu to get an idea of the different databases that you can access via Ugene:

Select Genbank and enter the accession number in the Resource ID box:

Click OK.
The sequence is loaded and visualized in the main window:

![]() You have to search for an accession number. You cannot use keywords like in the previous exercises e.g. gene and organism names...
You have to search for an accession number. You cannot use keywords like in the previous exercises e.g. gene and organism names...
However, Ugene allows to search Genbank based on gene, organism or author names via the Search NCBI Databases tool.
Retrieve sequences from Genbank based on gene and organism name
| Search for human ZEB1 sequences via Ugene. |
|---|
This opens the NCBI sequence search page
In the results list, you can select the record you want to download. Scroll down and select the record with ID = NM_001128128. Click Download:
 Click Ok to visualize the sequence in the main window. |
Downloaded sequences are visualized in the main window. Color-coded annotations and translations are added on top of the sequence. Annotations are fetched from the Genbank record, translations are added automatically by Ugene.

You can see a list of the annotations as they appear in the Genbank record by expanding NM_001128128 features in the bottom window:

When you search the record in Genbank you will see that the annotations in the record indeed correspond to the annotations in Ugene.
The annotations displayed in black like CDS, exon, polyA signal are shown, the annotations displayed in grey like STS are not shown on the figure. If you want to show them:
- Click the Annotations highlighting button at the right (red box on the figure)
- Click the annotation you want to visualize, e.g. STS (green box on the figure)
- Tick Show annotations of this type (blue box on the figure)

The STS are now visualized on the figure (purple box on the figure).
You can now use the sequence in Ugene for various analyses: BLAST, alignment, designing primers, restriction, ORF-finding... We will come back to these tools later.
![]() Even though you can search in some fields of Genbank records, Ugene doesn't give you the precision of searching that NCBI does
Even though you can search in some fields of Genbank records, Ugene doesn't give you the precision of searching that NCBI does
Exercise 6: Genbank searches on CLC Bio Main workbench (VIB only)
Exercise 7: genomic scaffold for Drosophila
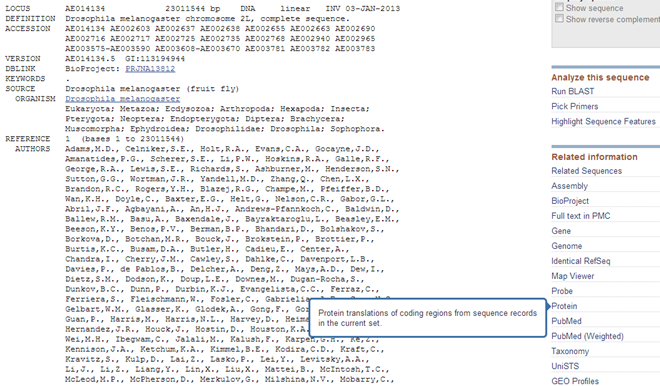
Search for the genomic scaffold AE014134 of Drosophila melanogaster. Go to the results in the Nucleotide database.
This is the sequence of a complete chromosome. You see that very long sequences such as these are by default not displayed at the bottom of the Nucleotide record (it would take too long to open the record in your browser, even as it is now it takes a very long time).
If you want to see the sequence in your browser you can customize the view.

| Download the protein translations to see the predicted proteins for this scaffold. |
|---|
This will generate a FASTA file containing the protein sequences in the Downloads folder of your computer. |