Install YASARA and its FoldX plugin
From BITS wiki
Part of Protein Structure Analysis training
Source: http://foldxyasara.switchlab.org/index.php/Installation_and_first_use
Installation
- Download and install YASARA
- YASARA View is the free version of YASARA and can be downloaded from the YASARA website after registration.
- You will receive an email with a download link. Please follow the installation procedure which is a simple extraction.
- Note down the location where YASARA has been extracted.
- The FoldX plugin for YASARA can be used in any version of YASARA: View, Model, Dynamics, Structure, Twinset.
- Download and install FoldX suite
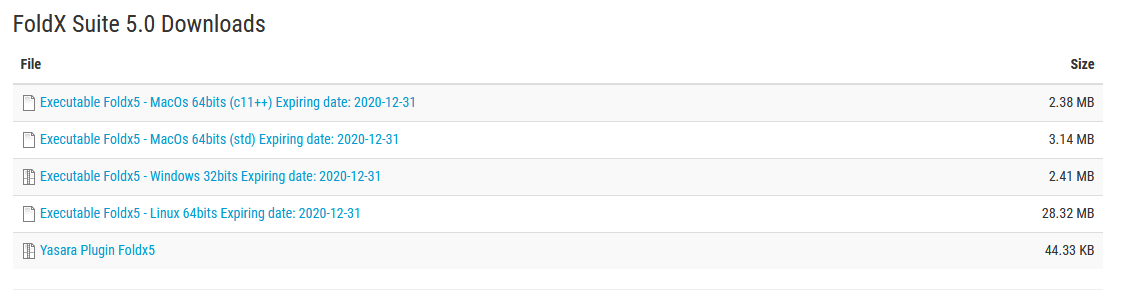
- The FoldX plugin for YASARA requires that you have FoldX installed on your computer as a 'stand alone' program. For compatibility purposes you need to download the latest version (FoldX 5) from http://foldxsuite.crg.es after free and simple registration for academic users.
- Please download the FoldX version 5 executable for your operating system and the Yasara plugin. Please scroll to the bottom of the page!
- Commercial users should contact [1] to acquire a license.
- Install FoldX by unzipping the file you just downloaded. This will unzip two files: an executable file and a file called 'rotabase.txt'. Both are necessary.
- Please note down the exact location of the two files. We will need this later on in YASARA.
- Using your file explorer or Finder, go to the folder where YASARA has been installed.
- The content of the plugin has to be extracted in the folder yasara/plg (or yasara\plg folder for Windows users) within the folder where YASARA is installed.
- After doing that you should have these 7 files (among many others) in your yasara/plg folder:
- foldx.py
- foldxaminoacids.txt
- foldxanalysecomplex.py
- foldxbuildmodel.py
- foldxplotoutput.py
- foldxrepair.py
- foldxstability.py
- Download and install Python
- On Linux and MacOSX machines Python is installed by default. Windows users can download Python from http://www.python.org/download or install it from within YASARA by clicking:
Help > Install program > Python
- On Linux and MacOSX machines Python is installed by default. Windows users can download Python from http://www.python.org/download or install it from within YASARA by clicking:
First use
Start YASARA and load a structure, such as 2AC0.sce. Now go to
Analyze > FoldX > Configure plugin
In the first browser window select the FoldX executable file.
As long as you don't change the location of those files or you don't overwrite the foldx.cnf file (be careful when unzipping a new version directly in the plg folder), this procedure has to be done only once.