Exercises on Protein Structure
Go to parent Basic bioinformatics concepts, databases and tools#Exercises_during_the_training
The PDB database
PDB (Protein Data Bank at http://www.rcsb.org/) is a database of experimentally determined 3D-structures of nucleic acids, proteins and their complexes. It is maintained by the Research Collaboratory for Structural Bioinformatics (RCSB). The structures have been determined by X-ray diffraction (on crystalized proteins) or NMR (on proteins in solution).
Exercise 1
Retrieve the structure for the human VH-1-related phosphatase (1VHR).
- go to http://www.rcsb.org/
- enter 1VHR in the search term box. Just like on the NCBI home page, it's a simple text box in which you can type PDB-IDs or keywords
- click Go button
More detailed search options, such as Advanced Search are available below the search box.

For structure information and search results, tabs can be used to toggle between different topics.

As said before PDB is highly redundant. In an attempt to remove the redundancy sets of highly similar structures were replaced by a single representative. In our case that representative is 3F81.A
| Check on the Structure Summary tab how many alpha helices and beta strands are present in this structure? |
|---|
| On the Structure Summary tab click the Show structure comparison... button next to see an outline of the secondary structure of this protein. At first sight you count 7 alpha helices (displayed in pink) and 5 beta strands (displayed in yellow).
 |
However, you need a more detailed overview of the secondary structure than what you see here.
| Check on the Sequence tab how many alpha helices and beta strands are present in this structure? |
|---|
|
You can find that on the Sequence tab in the Secondary Structure:DSSP section of Annotations - Details next to the figure. You see that they describe the presence of 9 alpha helices (the fifth consists of two separate alpha helices) and 7 beta strands since they include the two beta bridges (dark yellow triangles on the figure).
 |
VAST+, the enhanced version of VAST, short for Vector Alignment Search Tool, is a computer algorithm developed at NCBI and used to identify similar protein 3D structures by purely geometric criteria to identify distant homologs that cannot be recognized by sequence comparison.
| What is the most similar protein to 1VHR with respect to 3D structure according to VAST ? |
|---|
|
It's very easy to use, just enter the PDB ID in the search box and click Search. The first hit with PDB ID 4B04 is the most similar protein with respect to 3D structure. |
The ninth hit with PDB ID 3F81 is a copy of 1VHR in the NCBI structure database. You can see that this is a copy by clicking the ID on the VAST results page. This opens the record of 3F81 where you can see that 3F81 is VHR. The third hit with ID 1J4X is also a copy of VHR, in which a single mutation was introduced.
View the structure of the protein in YASARA. This software is already installed on the BITS laptops: click on the YASARA icon on the desktop to open the software.
![]() Before you start using YASARA, view the Working with YASARA movie to make you familiar with the user interface.
Before you start using YASARA, view the Working with YASARA movie to make you familiar with the user interface.
In YASARA go to: Help > Play help movie > General: Working with YASARA
YASARA can automatically fetch files from PDB:
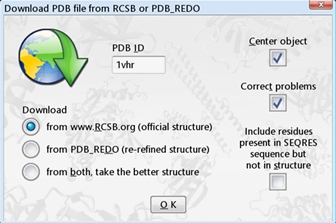
- File > Load > PDB file from Internet
- enter "1VHR" in the "PDB ID" box
- select "Download" "from www.RSCB.org (official structure)"
- click "OK"
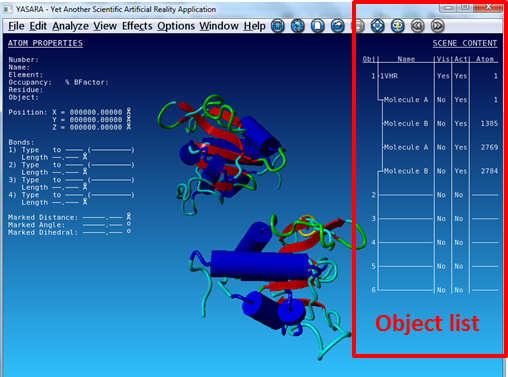
| Check the results of the VAST search in YASARA. First load 1VHR and 3F81 and align their A chains based on structure. |
|---|
First color the two objects in a different color:
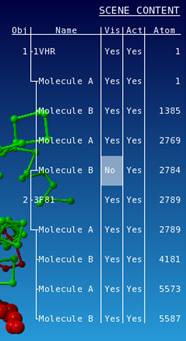
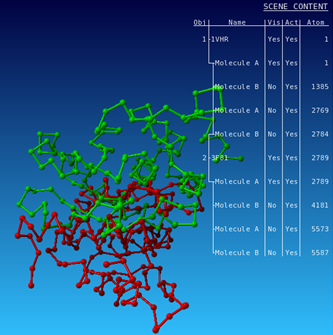
All molecules of 1VHR are now displayed in red. In the same way color 3F81 in green. 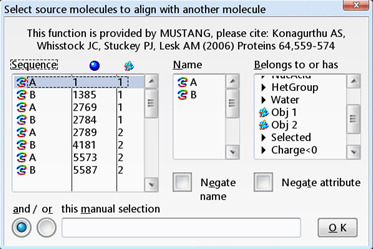 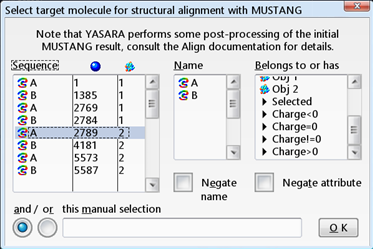 You see that they are indeed well aligned (RMD=0.58 Ä, 100% sequence identity). 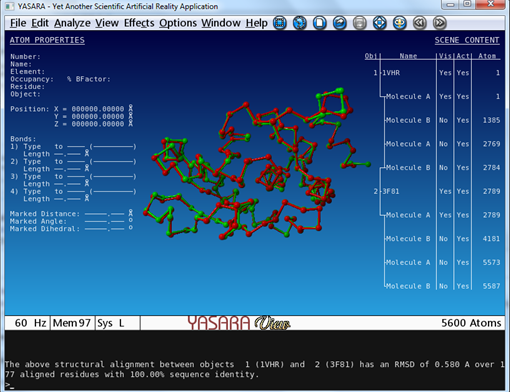 RMD is the root mean square distance between structurally equivalent atoms http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2374114/). |
Now do the same for 1VHR and 4B04: you will see that the alignment is not so nice.
Exercise 2
Find the structures of bHLH (basic helix-loop-helix) proteins in PDB. These proteins bind DNA and are involved in the regulation of gene expression.
You do not have to use a PDB ID to search PDB, you can also use keywords. Search PDB for "bhlh". In the left menu you will get an overview of ways to refine your query. Below you see a list of 26 PDB records that contain the word "bhlh".
We are looking for a yeast bHLH so use the Organism Refinements to filter the results for Saccharomyces cerevisiae bHLH structures. This takes you to the record of 1A0A.
Check the Structure Summary tab. The "Experimental Data Snapshot" section shows the method to solve the 3D structure, the resolution, and the overall quality of the structure (R free value).

Look at the detailed quality description with the sliders.
| Is this a good quality structure ? |
|---|
| No the sliders are at the right in the red section meaning that the majority of PDB structures and the majority of PDB X-ray structures with similar resolution score better on these criteria than this structure. Also the percentile values are very low. |
You can download the full quality report by clicking the figure.
| How many molecules are in this 3D structure ? |
|---|
In the "Macromolecules" section, you see that there are 4 molecules:
 |
| Download the PDB record for 1A0A onto your computer. |
|---|
|
At the top right of the Structure Summary tab you can click Download Files and select PDB Format
 |
This will save the file in your Downloads folder. Open the PDB file in a text editor like WordPad.
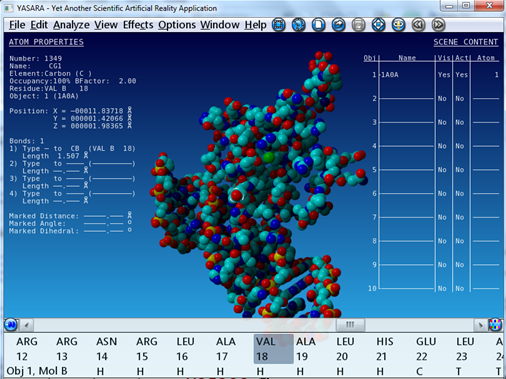
| What are the XYZ coordinates of the first atom in the structure ? |
|---|
|
The lines that start with the ATOM keyword contain the coordinates of the atoms: ATOM 1 O5' DC C 1 6.718 3.528 61.794 1.00 25.61 O The numbers 6.718, 3.528 and 61.794 are the XYZ coordinates in Angstrom of the first atom. |
| What kind of atom is the first atom ? |
|---|
You find the answer on the same line:
ATOM 1 O5' DC C 1 6.718 3.528 61.794 1.00 25.61 O The third column (O5') and the last column (O) tell you that the first atom is an oxygen atom. |
In YASARA, you can also load structures from a PDB file on your computer.
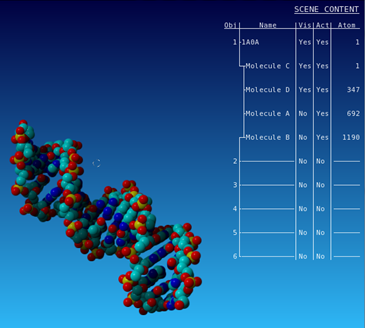
| Only display the DNA part of the structure |
|---|
|
| Visualize the secondary structure of the protein chains of the structure |
|---|
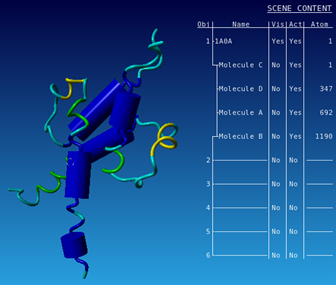
|
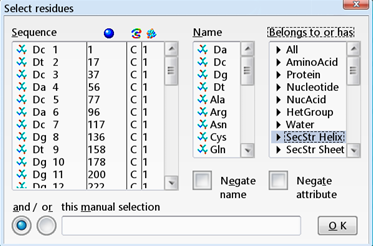
| Colour the alpha helices magenta |
|---|
This opens the "Select residues" window. |
Additional exercises
Visualizing_protein_structures_with_YASARA_exercises
Comparing_structures_exercises
Predicting_the_effect_of_mutations_on_protein_structure_exercises
Predicting_protein_structure_-_homology_modeling_exercises
Protein_ligand_interaction_-_from_small_molecule_to_protein_exercises