Archive for Module 2
Contents
FastA at the EMBL-EBI
Introduction of FastA
Pearson's FastA finds similar sequences in a database given a query sequence. FastA is older than the popular BLAST, it is slower, but sometimes finds things that BLAST does not find, especially for non-coding DNA (e.g. promotor sequences). We will use the FastA server of the EMBL-EBI, which has a richer collection of databanks. Go to http://www.ebi.ac.uk, then select the menu item "Tools"/"Similarity & Homology"/"FASTA".Error creating thumbnail: Unable to save thumbnail to destination
We will search with a fragment of the bacteriophage lambda genome containing the third right operator (OR3) and the right promoter (PR) to find similar sequences in the bacteriophage section of EMBL databank. (Note:running against the complete EMBL databank as compared to the bacteriophage section would take to much time for this exercise session - remember that fasta is rather slow).
| Get the sequence with accession number M25165 in fasta, download it and save as M25165.fasta. Try yourself first, click 'Show' for tips. |
|---|
| Since we have a universal accession number we can use all major portals. |
| You can use http://mrs.cmbi.ru.nl, search and download the sequence. |
| Or you can use http://www.ncbi.nlm.nih.gov/sites/entrez, search and download. |
| Or you can use http://www.ebi.ac.uk/, search and download. |
| |
By default the FASTA search page is set up for protein similarity search. Since we want to search nucleotide, we have to switch to the Nucleotide Databases' under 'Other types' in a box at the right side of the screen.
After that, we can select the database and our query sequence.
Exploration of the output:
- fasta program is run on the server of EBI with the parameters you provided. The "raw" output of the fasta program (text) has been postprocessed by the website into a nice layout. So now you are presented a list of sequences similar to your query sequence.
- First you might check the parameters by going to Submission details. Check if the database should be 'em_rel_std_phg'. Then go back to the summary page.
- In the actual fasta output you find the list of "hits", ordered by E-value. You can group them on the different columns by clicking on the header.
The first value you should check, is the E(xpect)-value, which is number of similar hits happen purely by chance (similar = similar length in a similar database). So 5.5E-13 means that once in 5.5*10^13 searches with a similar but random query and a similar but random database, such a hit will be found (by chance). So 5.5E-13 is a good E-value. If the databank sequence is longer, the E() values are better (lower) and the scores will be higher. This is because the chance to obtain a similar score for an unrelated sequence of same length is reduced.
The column called 'Length', 'identities' and 'positives' should explain themselves: the length of the hit found, the % of identical residues, and similar residues. In case we are dealing with nucleotides (as now), similar has been put equal to identical.
| Order the hits by DB:ID column. Check the ID's of the first two hits. What is their E-value? Which is most similar? |
|---|
| Click on the header of the DB:ID column to order by ID: 'EM_PH:AF069529' should appear on top. |
| The first two hits are actually recorded from the same sequence, AF069529 (out of the EM_PH database). The first hit has 1.3E-12 the second 0.17.
Error creating thumbnail: Unable to save thumbnail to destination
|
| What has happened here is that FastA has found two similar regions to our query. He has reported each hit separately. |
| So the first hit (1.3E-12) is the best hit. This is also apparent from the number of positives and identities. |
| Stay with the two hits of the task above: show the alignments of the two hits. How does the first two hits relate to each other? |
|---|
| First click on 'Clear selection' at the top of the table. Error creating thumbnail: Unable to save thumbnail to destination . Afterwards click on the first two hits.
|
| Click on Error creating thumbnail: Unable to save thumbnail to destination The alignments are shown in the results page. |
| For the first hit, the identity is close to 100%. But the second hit has a lower sequence similarity. |
| The query sequence is found back with very low similarity in the reverse complement (second hit). Interesting, and probably reflects secondary structures in the promotor region. |
- FASTA searches not only the query sequence but also its complement against the databank; this makes sense because we do not know in advance the orientation of eventual homologues in the databank. You can see this from the coordinates in the alignment for the first two shortest hits.
- To have nice graphical view of the results, click on Visual Output
Summary
- You might use Fasta when searching non-coding nucleotide sequences (UTR, promotor regions,...)
- Unfortunately, Fasta webinterface at EBI accepts only one sequence per submission
The NCBI VecScreen tool
Databanks sometimes contain errors. In this exercise we will prove that a human sequence from EMBL still contains some piece of cloning vector. We will use the VecScreen tool of the NCBI, which performs a BLAST search against the UniVec databank, which contains vector sequences from Genbank and some never published sequences from commercial sources, selected in such a way as to provide a nonredundant set.
You can start by taking a look at the file M34511 in the Nucleotide database of NCBI.
Then, go back to the NCBI BLAST home page and follow the link "vector contamination" in the "Specialized BLAST" list.
Error creating thumbnail: Unable to save thumbnail to destination
Copy-and-paste the accession number and "Run VecScreen". In the result page you will have to click "View report". The interpretation is rather obvious : the sequence is contaminated, although the reported exon falls outside the vector remnant. Note by the way that if you used a similar tool at the EBI, which runs a BLAST against EmVec, you would not have seen the contamination, because the vector sequence is not in EMBL.
BLAST: Mass spec peptide
| Solve the following. A peptide tag has been obtained from a mass spec experiment: GAGYGRALGGGSFGGLGMGFGGSPGGGSLGILSGNDGG. Can you see what happened? |
|---|
| BLASTing reveals similarity to human keratin. Most likely we are dealing with contamination. |
Pairwise comparisons
Nice graphical comparison of two sequences, using SIM6 and LALNVIEW
The SIM6 program of Huang and Miller calculates the best local alignment, and also a series of non-overlapping suboptimal alignments. Laurent Duret made a modified version of SIM that writes output in .lav format, suitable to be visualized with the graphical viewer LALNVIEW.
Go to the web site http://www.expasy.ch/tools/sim-prot.html. Although SIM can handle nucleic acids as well as proteins, the Web page at the SIB is configured only for proteins. Note: it is possible to download and compile the source code of the modified SIM and use it locally, also for nucleic acids.
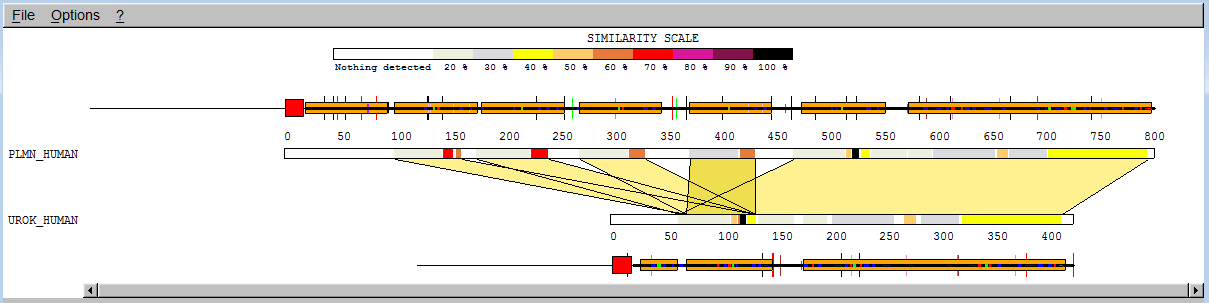
We have already mentioned the "kringle" domains. While the tissue plasminogen activator contains two "kringles", plasminogen contains five and the urokinase-type plasminogen activator contains only 1. Type plmn_human in the "AC or ID" box for "SEQUENCE 1" and urok_human in the "AC or ID" box for "SEQUENCE 2", then click "Submit". You will get a page with the classic SIM output.
Click on the "here" from "Click here to view these alignments..." and save the file 'view-laln-file.pl' (or whatever you wish to rename it). Start LALNVIEW on your PC: you can find it under "All programs". Select "File"-->"Open" the file.
You see 2 alignments with a score of more than 120 (BLOSUM62) represented by coloured bars. You can click on any part of the coloured bars to adjust the alignment of the bars. Only similar portions with a score higher than 120 are displayed. You can lower this score by replacing 120.0 in the "Similarity Score Threshold" box by 50.0 and press <enter>.
You will then see 5 alignments. To get more information on a portion of the sequence, note the additional bars above and below the upper and down sequence bar. Clicking on those will reveal the name of the domain and motifs. Note however that this information has simply been extracted from the annotation that accompanies the sequences in the SwissProt databank, it has not been computed.
We can see that the first 4 correspond to an alignment of a "kringle" of plasminogen with the unique "kringle" of urokinase, the 5th alignment encompasses the last "kringle" of plasminogen as well as all the motifs that follow. If you click on a coloured bar, the alignment will appear in a second window. If you set the threshold to 0 you will see even more alignments, up to 20 ; the reason is that SIM cannot evaluate significance, it continues to compute non-overlapping alignments with ever decreasing scores until no alignment with score higher than 0 can be found or until a limit (here set to 20) is reached.
There exists also a SIM2 program that computes local and suboptimal alignments using the fastA algorithm (and is in fact identical to the lfasta program from the fastA suite); SIM3 and the improved version SIM4 [1] are designed to align a gene with a mRNA, with a provision for finding splice sites.
The graphical way of detecting similarity: dotplots
We will use Dotlet to investigate the usefulness of dotplots. This is a JAVA program, accessible on http://myhits.isb-sib.ch/cgi-bin/dotlet. A dotplot program such as this creates a visual representation of the sequence similarity of two sequences. In the folder C:\data-bioinfo, you will found the sequences X62302 and the M24173. One is a gene sequence, the other cDNA. You can search more details from these sequences by using the accession number.
Let's put those two sequences into DotLet.
Using the input button, you can import two sequences: this must be done by copy-paste. For every sequence, you have to click this button.Select from the second and third dropdown box respectively the two sequences. Click compute with the default settings.
The left graph shows you the dotplot graph, the right graph shows you the score distribution. Above the right graph, the parameters are summarized.
On the dotplot graph, one sequence is placed on the vertical axis, another on the horizontal axis. If the residue on the Y-axis is identical to the X-axis, a white dot is drawn. If the residue are different, a black dot is drawn. The complete graph looks as a random collection of black and white dots. But where the sequences are similar, white diagonal lines are visible. To zoom out, change the last button at the top, from 1:1 to 1:7. The button before that button, which is set by default on 15, you can set this to 43. This is the window size: to draw a white dot, if set to 15, the 7 residues before and the 7 residues after that position needs to be identical.
Click compute with the new settings, and you will see a more meaningful plot.
The longer sequence (gene) is on the X-axis, the shorter (cDNA) on the Y-axis. In the dotplot window there are crossed blue lines, you can drag the crosspoint with the mouse, causing the text in the sequence window at the bottom to follow, according to the position of the cross. You can move one base at the time by using the arrow keys. You can estimate the positions of the three exons (and by consequence of the two introns) by projecting the diagonals of similarity on the horizontal axis. The blue lines are a great help for doing this with precision.
| Why are the diagonals in the dotplot not a 100% black line and often have small gaps? |
|---|
| The gray scale at the right shows you what the scale means. A window can have score 0: none of the residues in the window correspond. With a window of 43, the maximum score is 43: all residues in the window overlap. Now this occur rarely: you can set the gray ramp to help you visualise these scores. |
| By dragging the ramp to the right, only higher scores will appear as white dots. Lower (hence less similarity) will be blackened out. |
Dotplot of protein sequences
The human tissue plasminogen activator precursor (in UniProt/SwissProt with ID TPA_HUMAN and AC P00750) contains two "kringle" domains. "kringles" are protein domains (protein sequence parts that are shared between different proteins - see next module) that have a typical structure and are involved in the interaction of certain blood proteins with each other and with cell membranes. We will see protein domains in the next part: sequence analysis. But for now: to know more about this protein domain, you may want to check this site.
You can find the file TPA_HUMAN.fasta in the C:\data-bioinfo folder, to solve the next exercise.
| Make a dotplot from TPA_HUMAN.fasta (acc nr: P00750 - Tissue-type plasminogen activator) against itself. What do you see? |
|---|
| Use Dotlet and input the TPA_HUMAN.fasta |
| We see on the plot the presence of a full diagonal, which is normal since a sequence is always 100 % similar to itself. |
| The segments outside the main diagonal reveal a repeat: these are the two kringle domains. You can find the position of the two "kringle" domains by projecting a segment on the two axes. |
| You can position the blue crosshair on one of the segments, so as to reveal in the sequence window an alignment between the two "kringles" ? Note how identical amino acid pairs are indicated in cyan and "similar" amino acid pairs (positive score in BLOSUM62 table) in dark blue. |
| Error creating thumbnail: Unable to save thumbnail to destination
|
The sequences that will save your scientific life
An experiment of yours concludes that two proteins are similarly regulated. So you get the sequences of those proteins. Funny though, at first sight, you cannot find a similarity between the proteins. Is this a mistake?
>Similarly_regulated_protein1 KGMYYKNEHFTDKCSGNPFHGQSCAFHNIWIPRLFGPQEKVEPASCTHCGDGLYSAPQAL DLEEQTAWTSGPWTCRKYVPVCNKEWLNFKDIEQCGCGGHKLPYYVMSSMMWTPGTTHVK >Similarly_regulated_protein2 GEHFHPTNDDMCGCGGHKLWFQEQHGIFALKSPEDYMDFGQMQSTNTIDWIALNVCWEYI ECFTLPTCTAWWKKGSLCLFDHFCIKCWMDHIWLNHIMMDNCRNNYRSHSMGINLMVFLW
| Throw all your techniques on the table to look at those sequences. |
|---|
Error creating thumbnail: Unable to save thumbnail to destination
|
########################################
# Program: needle
# Rundate: Thu 10 Feb 2011 13:03:00
# Commandline: needle
# [-asequence] /ebi/extserv/old-work/needle-20110210-1302585701.input.1
# [-bsequence] /ebi/extserv/old-work/needle-20110210-1302585701.input.2
# -outfile /ebi/extserv/old-work/needle-20110210-1302585701.output
# -gapopen 10.0
# -gapextend 5.0
# -datafile EBLOSUM62
# -sprotein1
# -sprotein2
# -auto
# Align_format: srspair
# Report_file: /ebi/extserv/old-work/needle-20110210-1302585701.output
########################################
#=======================================
#
# Aligned_sequences: 2
# 1: random
# 2: random
# Matrix: EBLOSUM62
# Gap_penalty: 10.0
# Extend_penalty: 5.0
#
# Length: 203
# Identity: 9/203 ( 4.4%)
# Similarity: 14/203 ( 6.9%)
# Gaps: 166/203 (81.8%)
# Score: 36.0
#
#
#=======================================
random 1 KGMYYKNEHFTDKCSGNPFHGQSCAFHNIWIPRLFGPQEKVEPASCTHCG 50
random 0 -------------------------------------------------- 0
random 51 DGLYSAPQALDLEEQTAWTSGPWTCRKYVPVCNKEWLNFKDIEQCGCGGH 100
.|..:..:.:.||||||
random 1 ---------------------------------GEHFHPTNDDMCGCGGH 17
random 101 KLPYYVMSSMMWTPGTTHVK------------------------------ 120
||.:.....:..........
random 18 KLWFQEQHGIFALKSPEDYMDFGQMQSTNTIDWIALNVCWEYIECFTLPT 67
random 120 -------------------------------------------------- 120
random 68 CTAWWKKGSLCLFDHFCIKCWMDHIWLNHIMMDNCRNNYRSHSMGINLMV 117
random 120 --- 120
random 118 FLW 120
|
Alignment editors: GeneDoc
Start GeneDoc and open the file ADHs.msf using the "File"/"Open" menu. You will see that GeneDoc has displayed the alignment using different shades of grey to indicate the degree of conservation. GeneDoc can present the alignment in a variety of ways and is very suited to prepare alignments for publication. To edit your alignment, put Genedoc in "Grab and Drag" mode: do this by selecting with the mouse the "Arrange"/"Grab and Drag Sequences" menu option, but also by clicking on the most left icon
Alignment editors: BioEdit
Another good free editor (but not well maintained) I want you to know is BioEdit. It has alignment algorithms build in. It can read a variety of sequence and alignment formats, can color the sequence on different criteria and is very convenient for editing. In addition, you can ask statistics of the alignment, such as consensus sequence, similarity and identity matrices etc.
| If not already present on your computer, install Bioedit (Windows only - other OSes may choose to have a look at Jalview) |
|---|
Use BioEdit to align the 8 plant Histone H1 sequences in fasta format you have retrieved during previous exercises. First open the fasta file, and next, choose 'accessory application' -> 'ClustalW multiple alignment'
Error creating thumbnail: Unable to save thumbnail to destination
Once the alignment is completed, you get the colourful results which can edited it further.
the above selector you can switch between 'viewing' and 'editing' mode: in the Select/slide mode, the mouse can be used to select sequence parts. Once selected, sequences can be copy-pasted. In the edit mode, the mouse acts as a cursor and sequences can edited. In the Grab/drag mode, the mouse can be used to drag a sequence forward in the alignment. However, BioEdit is not being properly maintained anymore, so be careful for crashes and save your work regularly.
The alignment can be coloured according to the different similarity matrices. Go to 'Alignment' -> 'Similarity Matrix (for aligment and shading)'. Default, the Identity matrix is selected. You can select other, for example, PAM40.
| Predict what will happen with the colouring if you select 'higher' PAM matrices (PAM80, PAM120). And check it changing the colouring. |
|---|
| Recapitulation: PAM tables are derived from PAM1, a scoring matrix derived from highly similar sequences (differing 1%). Higher PAM numbers are multiplications of the PAM1 matrix. |
| Hence, the PAM1 matrix gives more weight to highly similar residues, the higher matrices give relatively more weight to less similar residues. |
| So, when colouring with higher PAM matrices, columns containing more dissimilar residues will get also coloured. |
| You can check this easily: PAM40 colouring Error creating thumbnail: Unable to save thumbnail to destination PAM120 colouring Error creating thumbnail: Unable to save thumbnail to destination
|
Note that at the C-terminal end a lot of prolines appear (embedded in basic residues as R and K). We will demonstrate the following principle in Bioedit.
| How can you make sure all prolines stay as much as possible aligned in the alignment? |
|---|
| Let's think in reverse: if prolines need to stay aligned, the optimal path when determining the pairwise alignment should run through P-P positions. So achieve this, a P-P match should receive such a score that it contributes to a large extent to the final score. Doesn't this sounds logical? |
| How do we implement this, so we can align all prolines in the C-terminal part? It is indeed all about changing the substitution matrix used for scoring the alignment. |
| In the installation folder of BioEdit, a folder 'tables' can be found. Substitution matrices can be found here. We will modify the BLOSUM62 matrix to our needs. First copy this matrix to a new file with the name 'matrix_proline.txt'. Open the matrix in Wordpad, and change the value for a 'change from P to P' to 100. Save and close Wordpad. Now, rerun ClustalW of the histones with BioEdit. In the first window, enter as a parameter "/MATRIX=C:\BioEdit\tables\matrix_proline.txt". |
|
Error creating thumbnail: Unable to save thumbnail to destination Leave the remaining parameters as default. See what happens when you run the alignment? |
Error creating thumbnail: Unable to save thumbnail to destination |
Go to 'File' -> 'Graphic View' for a graphical view of the alignment. You can set numerous shading options. To export the file, export it as 'rich text format' (rtf). This can be opened in Word and be included in an article.
Summary
Pairwise similarity detection can be done with graphical methods (dotplots), optimal methods (water and needle), which take a long time to compute, and heuristic methods (BLAST), which are fast, but do not guarantee you the optimal result.
Your gene of interest
Search the position in the genome using BLAT and extract the sequence
For your gene of interest, use the genomic nucleotide sequence from RefSeq to do a BLAT search to the genome, using the BLAT search portal. BLAT can align a nucleotide sequence to genome if the sequence similarity is high enough (>80%).
The results-page lists portions of the genome with high similarity. Don't pick the 100% similarity, but look for a hit with lower percentage similarity. How similar is this portion of the genome? How long is the hit? Can you extract the nucleotide sequence of the genome of that hit in fasta format?
Hint: click on the second entry in the left panel
Align the genomic sequence
Align the genomic sequence with all other sequences that you have retrieved in Module 1 exercises. Use any clustal omega (e.g. from ebi http://www.ebi.ac.uk/).
- How divergent are the sequences from that genomic sequence part?
- Can you generate a logo from the most similar part?
Go to parent Basic bioinformatics concepts, databases and tools#Exercises_during_the_training
Go to parent Basic bioinformatics concepts, databases and tools#Exercises_during_the_training