Pairwise sequence alignments
Go to parent Basic bioinformatics concepts, databases and tools#Exercises_during_the_training
Pairwise sequence comparison is widely used in sequence analysis. We can discern three approaches:
Contents
Dynamic programming: global and local approaches
An optimal alignment can be calculated, if you provide a scoring scheme (for matches and mismatches) and a way to deal with gaps (gap penalty and gap extension scores). Two popular algorithms for this differ in the way they align the sequences: either they make the ends of the sequences match (global alignment) or they don't require the ends of the sequences to match (local alignment).
Exercise 1: MYOD1
We will compare the MyoD1 proteins of several species: human, mouse and fruitfly. We can compare them pairwise. The MyoD1 gene plays a role in the differentiation of myoblasts and its protein product contains a DNA-binding helix-loop-helix motif (see module 3).
Go to the EBI's pairwise alignment tools website. This page gives you access to the EMBOSS programs needle and water (which implement, as you might guess, the Needleman-Wunsch and the Smith-Waterman algorithms).
Use needle (global alignment) to align the human and mouse MyoD1.
- Upload or paste human MYOD1 and mouse MYOD1 into the upload boxes
For the moment, we will go with the default settings for scoring matrix, gap opening penalty and gap extension penalty. - Click Submit
The results are what we expected: the two homologous sequences are nicely aligned over the complete length.
Important parameters can be found in the header of the file:
- The scoring scheme that was used (substitution matrix, gap opening penalty, gap extension penalty)
- The length, number of identical and similar amino acids, number of inserted gaps, and similarity score of the obtained alignment
The BLOSUM62 matrix is used, although EMBOSS calls it EBLOSUM62. EMBOSS considers two amino acids similar if their corresponding value in the BLOSUM62 matrix is positive (identical amino acids are also counted as similar).
Below the header the report shows the best alignment that it could make with our parameters settings. You see that this alignment was not really difficult.
EMBOSS uses an output style where aligned amino acids are connected by: | if they are identical, : if they are positive and . otherwise.
| What is the outcome of the water algorithm with default parameters to align the same sequences ? |
|---|
| Water calculates the optimal local alignment. Proceed in the same way as for the needle algorithm. You see that you get the same alignment. |
Of course, it gets more challenging with more divergent sequences. Download the fruitfly MyoD1 sequence.
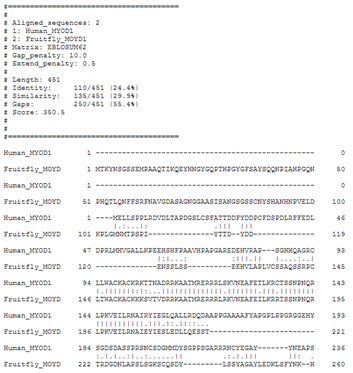
| What do you think of the needle alignment with default parameters of the human and the fruitfly MyoD protein ? |
|---|
|
The resulting alignment should be the best alignment, but you see that a lot of gaps are introduced. Of course, this is because the best alignment depends on the parameters! |
Let's try to get a more compact alignment by fine-tuning the parameters.
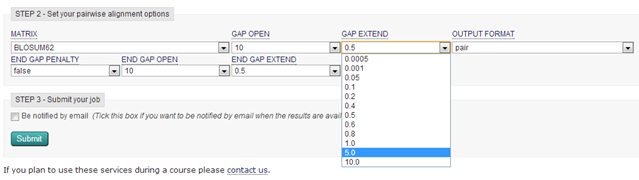
| What happens when you increase the gap extension penalty to 5 ? |
|---|
| To reduce the number of gaps, you have to adjust the gap opening and extension penalties. This is a case of trial and error. Of course you have to increase the gap penalties to prevent gaps being introduced the whole time. The resulting alignment is much nicer! A limited number of short gaps are now introduced. |
From the global alignments, it is clear that the central portion of the sequence aligns best.
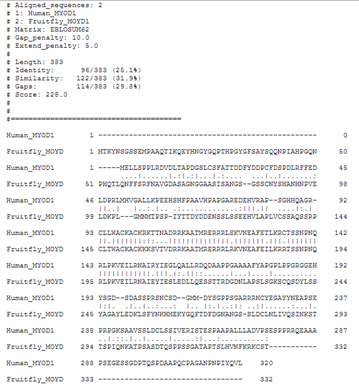
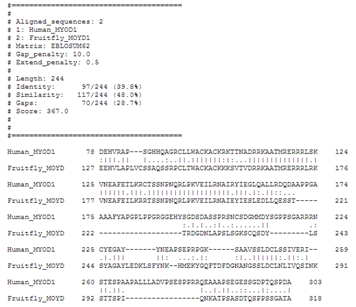
Let's see what water (local alignment) tells us about these sequences.
So, local alignments can help you to align only the best matching portions of a sequence.
The needle and water algorithms can also be used to align DNA molecules: click Nucleotide on EBI's pairwise alignment tools website. Aligning DNA causes the programs to use a +5/-4 match/mismatch scoring scheme (EDNAFULL matrix of EMBOSS).
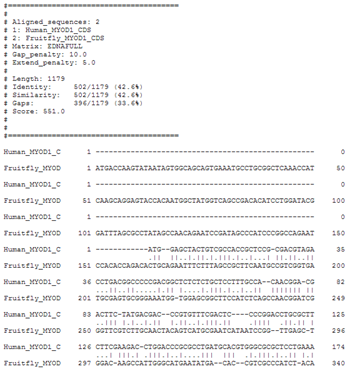
Perform a global alignment of the human and the fruitfly MyoD1 CDS.
Reminiscing the results of the global protein sequence alignment, it might be a good idea to use a gap extension penalty of 5 again.
One more important thing: local alignment will align the best aligning portion of your sequence. Sometimes, multiple similar regions exist in your sequences, but local alignment will only align the best and/or longest match, hiding the other similar regions...
Keep this in mind! Always try multiple tools and multiple parameter settings for aligning your sequences.
The graphical way of detecting similarity: dotplots
Exercise 1: mouse embryonic zeta-globin
We will use Ugene to investigate the usefulness of dotplots.
| Open the mouse zeta-globin mRNA (NM_010405) and gene (X62302) sequence in Ugene. |
|---|
| Upload the sequences in Ugene via Access remote database in the File menu using the Genbank IDs. |
We want to see the location of the introns so we need a dot plot with a small window. The window size is called minimum repeat length in Ugene: we will use 10 nt.
| Create the dot plot to compare the two sequences (use the Actions menu). |
|---|
|
To create a dot plot:
 This opens the window containing the parameters of the dot plot:
 Using these settings, the program will draw a dot if a 10-letter word in the mRNA sequence is identical to the gene sequence. |
You can minimize the windows with the sequences using the buttons indicated in the figure.

| Where are the introns located ? |
|---|
|
 Where the two sequences are identical, diagonal lines are visible. In this way it's very easy to see the location of the two introns (red on the figure). The first exon is located between around 600 and 1400, whereas the second exon is much smaller: between 1600 and 1700 in the gene sequence. |
| Check in the gene annotation whether these estimates are correct. |
|---|
|
You can find the gene annotation by expanding X62302 features in the bottom window.
 You see that our estimates are indeed correct. |
On the dot plot you see that a region near the end of the mRNA sequence is repeated in the gene sequence (green on the figure):

| Identify the sequence of the repeated region. |
|---|
|
You can find the sequence and location of this region by clicking it on the dot plot:
 If you click the region on the dot plot, it is automatically visualized on the sequence plot:
 |
You see that you can easily identify introns and exons using dot plots of mRNA and corresponding gene sequences.
Exercise 2: human TPA
Human apolipoprotein A (UniProtKB/Swiss-Prot ID: P08519) contains 38 kringle domains. Kringles are protein domains (see later in this module) that have a typical structure and are involved in the interaction between blood proteins and cell membranes.
Create a dot plot of the protein against itself in the same way as in the previous exercise (word size = 10; threshold = 100%)
| Make the dotplot. |
|---|
|
Load the sequence from UniProtKB/Swiss-Prot and create the plot.
 |
This creates a rather spectacular plot with a perfect diagonal line as we expect from aligning a sequence to itself and a very high number of repeats.
By clicking a repeat near the upper left corner of the plot we can see the location and the sequence of the repeat. When you open the human apolipoprotein A sequence in a text editor and you look for this sequence, you see that it is indeed repeated many times just as the dot plot shows you:

| Is this sequence part of the kringle domain? |
|---|
|
When you look in the annotation you see that this repeat (e.g. located from 33-58, 147-172...) indeed corresponds to a part of the kringle domain:
 |
So we have visualized the kringle but the plot looks a bit messy especially at the right and bottom edges.
| Check the annotation to check how long the kringle domains are |
|---|
|
When you look in the annotation you see that kringle domains are around 110 amino acids long.
 |
When you look in the sequence you see that kringle domains are very similar but not entirely identical so it might be a good idea to choose a lower threshold.

| Make a new dot plot with word size = 100 and threshold = 95%. |
|---|
| You see the first 29 repeats of the kringle domain:
 |
Apparently, the kringles at the end of the sequence are less conserved, which is indeed true when you look at the sequence:

Exercise 3: Compare chromosomes
One of the most useful applications of dot plots is that they give an easily interpretable representation of the alignment of very long sequences like full chromosomes.
We will compare the complete RefSeq sequence of Chromosome I and Chromosome VIII of Saccharomyces cerevisiae S288c.
| Open the sequences in Ugene via Search NCBI Genbank in the File menu. |
|---|
 Select the RefSeq sequence to download. |
we will create a dot plot of the two sequences using word size = 50 and threshold = 100%.
| Make the dot plot. Make sure Search for inverted repeats is selected! |
|---|
|
 |
| How many identical regions can you find ? |
|---|
 It is clear that there are 4 long identical stretches between the chromosomes (and also some smaller identical sequences) representing sequences from one chromosome that have been duplicated during evolution and that were inserted into the other chromosome. |
Three of these inserted sequences lie parallel to the main diagonal. The fourth one is perpendicular to the main diagonal,
| Why is the latter insertion different ? |
|---|
 Three of these inserted sequences lie parallel to the main diagonal (red on figure), indicating identical regions located in the same reading direction on both chromosomes. This means they are located on the same strand on both chromosomes. The fourth one (green on figure) is perpendicular to the main diagonal, meaning that this region on the reverse complement strand of one chromosome is identical to a region on the plus strand of the other chromosome. |