Exercises on Gene Ontology, protein structure and other non-sequence data
Go to parent Basic bioinformatics concepts, databases and tools#Exercises_during_the_training
Version 5 March 2011 by JJ
Version 4 Nov 2011 by JJ
Version 3 Feb 2011 by Joachim Jacob
Version 2 by Joachim Jacob
Version 1: Guy Bottu
Contents
- 1 Basic exercises
- 2 Additional exercises
Basic exercises
Following exercises are recommended to complete. If you have done these exercises, additional exercises can be found at the bottom of this page.
We are reaching the stage in which we are integrating many different databases: we will discuss the most important (personal opinion) in this section. The goal is that you understand most bioinformatics concepts and sources that you encounter online and in papers.
Gene Ontology
The Gene Ontology is the fruit of a collaboration between managers of several databanks. It is used a lot to fetch relevant genes and to interpret high-throughput data. We will look what information the GO database contains. The purpose of GO is to agree on standardized keywords. A collection of such standardized keywords is called "ontology" or "controlled vocabulary" and is becoming very important in this high-throughput era. The GO terms are widely referenced to by numerous databanks (EMBL-EBI, UniProt, ...). Besides being useful standardizing the annotation of the databanks (and hence making searching easier), GO is also useful for the interpretation of high throughput experiments. In short, by looking at the GO terms associated with the genes on a gene-list, overrepresented GO terms can be detected, and the biology behind the list might be inferred.
Searching genes based on ontology
Go to http://www.geneontology.org. In the "Search for genes, proteins or GO terms using AmiGO:" box type cytochrome c and "GO!". You will get a list of gene products with each time a link pointing to information about the gene product and a link with "associations": these are the GO terms that have been associated with that gene.
One of the entries you see close to the top should be
Click on "cyc1", note that you have a link to the Schizosaccharomyces pombe entry in the Taxonomy databank of the NCBI and another to the cyc1 entry in the GeneDB, which is one of the members of the GO consortium. Go back and click on "8 associations", you will get a list with the GO terms that have been associated with the gene cyc1 from S. pombe as it is in the GeneDB. Note that there are 3 "base" ontologies : biological process, cellular component and molecular function. Click on "mitochondrial electron transport, ubiquinol to cytochrome c". You will get an overview of the place of the term "mitochondrial electron transport, ubiquinol to cytochrome c" in the GO hierarchy.
Also in the GO databank, each entry has an GO accession number: you can use it to search databases for cross-references to GO. As an example, go to http://mrs.cmbi.ru.nl, select databank "Uniprot KB" and search for "dr:0042775". This will search for the GO accession number '0042775' in the database reference fields, corresponding to GO:0042775.
GO not only references genes and gene products, but also domains (InterPro database).
| Get some more information about the terms assigned to the retinol-binding domain (IPR002449). |
|---|
| Browse the IPR domain with the amigo browser Tip: you can also search at Interpro and click through to GO website or EBI's Quick GO http://www.ebi.ac.uk/QuickGO/. |
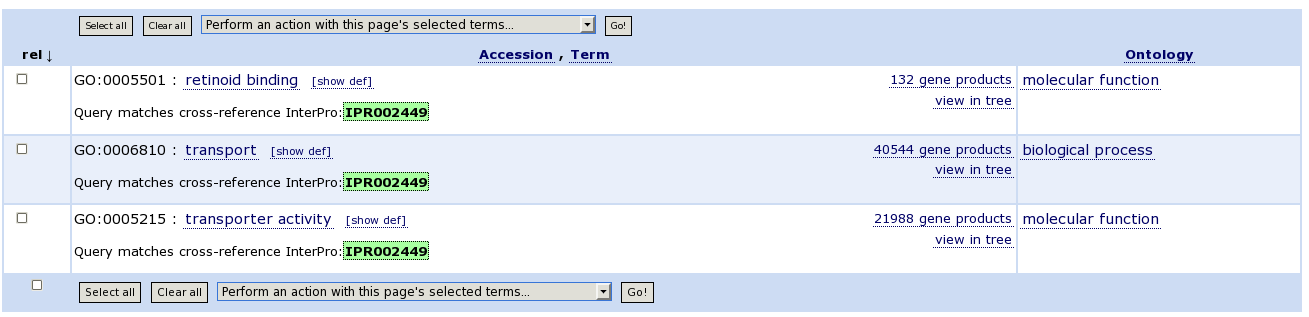
| On the AMIGO browser, search for the domain in the GO terms |
| You can find IPR associated to one biological process term, and 2 molecular function terms. These three term descriptions should give you already a nice picture of what IPR002449 is involved in. |
| To visualize its parent terms, we can view the tree of all GO terms subsequently. BP: establishment of localisation, MF: isoprenoid binding and molecular function (the 'root'!). So, a catalogued term can reside on different 'levels' in different ontologies. |
Entrez
By giving some examples, we will explore what (non-sequence) information you can retrieve from the NCBI databases using Entrez, in the meanwhile explaining some terms.
Searching some biological information about prion proteins
Prions are proteins which can change its structure in such way that other prions are stimulated to adopt the same conformational change. Let's explore them through entrez.
Go to http://www.ncbi.nlm.nih.gov/sites/gquery. Enter 'Prion' in the search box.
Let get more information about the related diseases, the structure and the expression profiles. Knowledge about which database to search for is indispensable for this.
PubMed with MeSH terms
What diseases are prions involved in?
You can look for a good review article for this: click the pubmed link which brings you to PubMed, the online portal of MedLine.
You may notice that PubMed found a MeSH (medical subject heading) term: a controlled vocabulary (just as Gene Ontology is) for medical terms (see in right column below). You can search PubMed for these terms to make sure you retrieve what you are looking for (More info on MeSh).
Other useful links can be found at the right side of the screen. For example, to find only reviews:
Diseases from OMIM db
Go back to the main Entrez results page: clicking on the OMIM database will show you genes which are annotated as prions and are involved in diseases. It provides more information on how prions disrupt the normal functioning.
Gene expression profiles from GEO
Go back to the main Entrez results page. Click on GEO profile, the database that contains expression information from micro array experiments.
GEO searches the search term in the annotation of genes, it then retrieves the appropriate related microarray probes. For the experiments for which these probes are significantly different from the background (hence, up or downregulated in the experiment), GEO displays the results, with most significant on top.
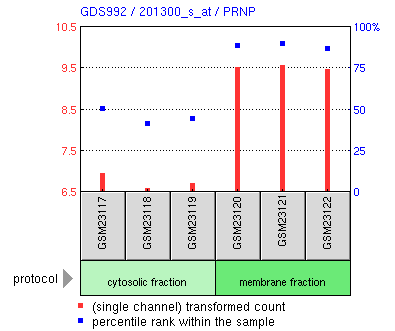
From this prion search in GEO, we can see that apparently the PNRP gene (the same gene as seen in OMIM) is more abundantly present in the membrane fraction compared to cytosolic fraction. To know more about conditions in which PNRP is depleted or enriched, you can search PNRP in the search box of GEO profiles.
GEO datasets (GEO identifier starting with 'GDS') are curated microarray experiments. Some analysis tools are available for these sets: for example to retrieve expression values in certain conditions.
Entrez and NCBI help pages
There are more databases at NCBI which can be search with Entrez, but we won't cover them all here. To help you browse their tables, NCBI has provided very nice help pages, which you should check before using any resource:
KEGG, a resource for pathway information
NCBI provides some nice integration of various resources, but the pathway information they get from KEGG. KEGG collects pathway information and corresponding genes and chemicals associated to pathways, in their subdivision KEGG pathways.
Let's have a look at KEGG: go to http://www.genome.jp/kegg/. Search for the term 'prion'. You should have one KEGG Pathway result: map05020
Prion diseases.
You can click on the map to have a nice big clickable map, ready for you to investigate. Have a look at the displaying map. Other linked pathways can be readily accessed, and orthologous genes can be fetched for any step in the pathway.
A complementary resource for pathway information is Reactome at EBI: http://www.reactome.org/ReactomeGWT/entrypoint.html. If you want nice graphics of pathways, you might want to check Biocarta Pathways.
Biograph, searching connections in biological data
When you want to know whether data exists that sustains connections between two biological entities, you might want to check Biograph.be. Biograph has mined literature for you, explored interaction databases, and more heterogeneous information sources: it integrates all those sources, allowing you to find connection between the information.
We will look whether their exists a connection in all biological data between proteins containing the chorismate domain and the chemical salicylic acid. Go to http://www.biograph.be.
At the right, you find text fields where you can enter any biological information: genes, pathways, diseases, chemicals,... Biograph will try to connect those entities for you based on available information. Indeed, it is a kind of knowledge generator... Type in 'chorismate mutase' in the left text field, and 'salicylic acid' in the right text field. Click on 'Find links'.

First biograph needs certainty over what you mean with chorismate mutase. Indicate the domain option from the list on the right. Then, click find functional links.
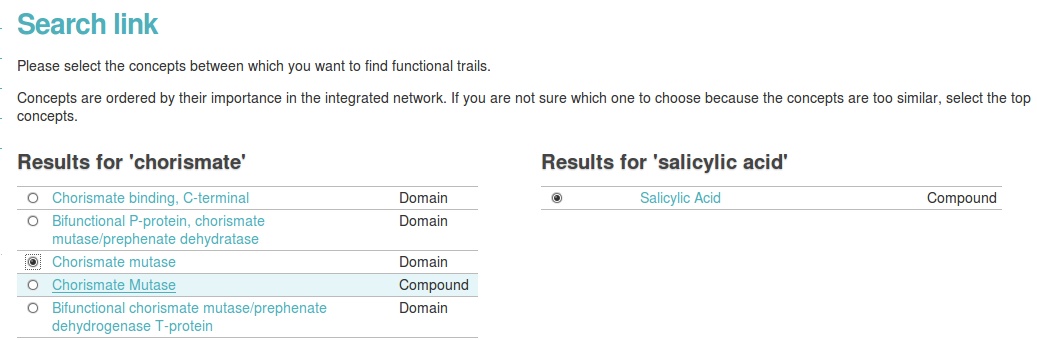
Finally, you enter a page with the title Relations between 'Chorismate mutase' and 'Salicylic Acid'. You find more info about the search, and a nice overview graph, displaying the path between two entities. This is a lot of information! But it can be potentially valuable. You can check the links by clicking on the edges or the nodes to view the corresponding evidence data.
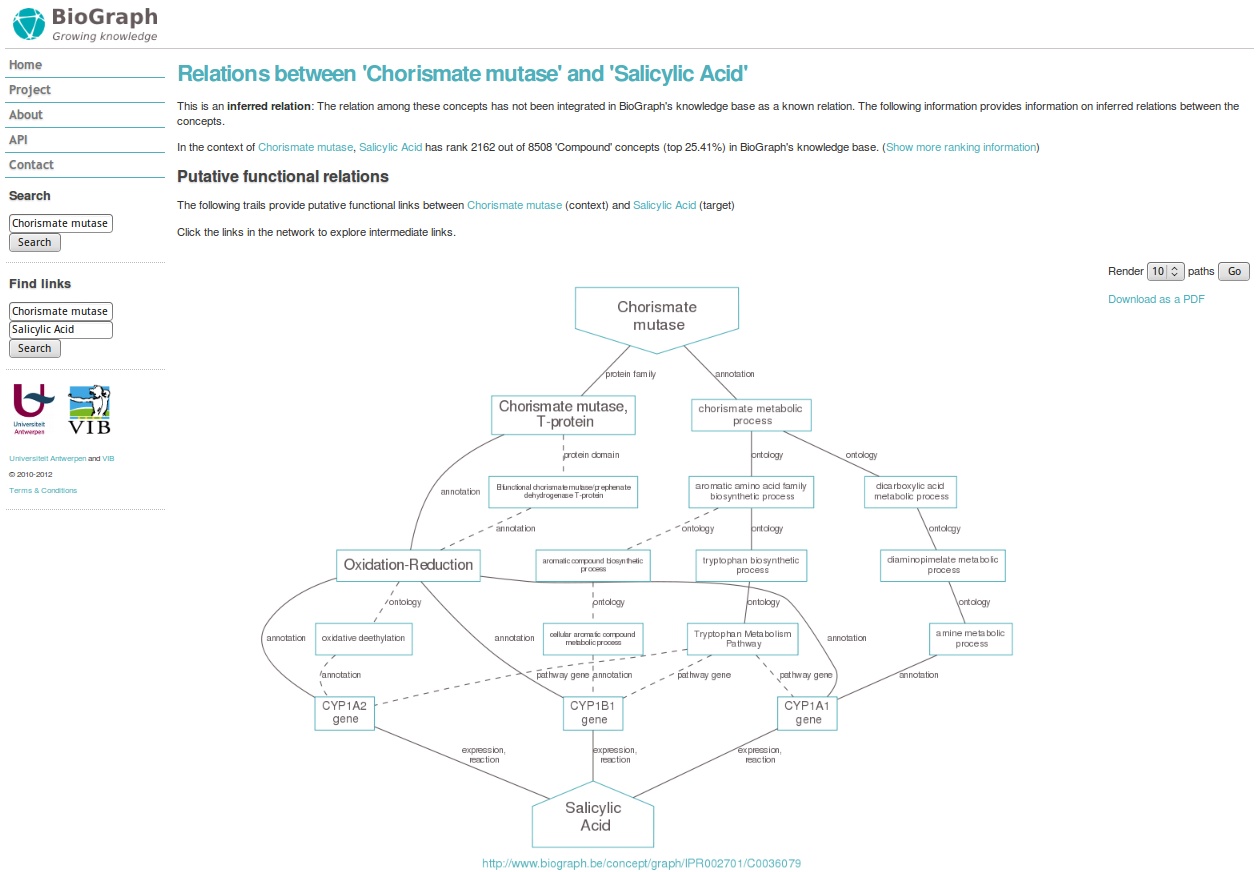
Try Biograph out with some terms or biological entities that you are interested in!
Integrating different information sources for your gene of interest using BioGPS
All biological information is spread over numerous webportals of the corresponding databases. It is cumbersome to browse around to all those different pages: this is where BioGPS comes into play.
Access BioGPS by going to http://biogps.org
It is a gene based portal: meaning you provide a gene name, or symbol, or a corresponding accession number, and all information related to your entry is retrieved.
Let's try it: type in 'TTR' into the search box at the middle, and click ![]()
The output is displayed in a new window, as shown below:
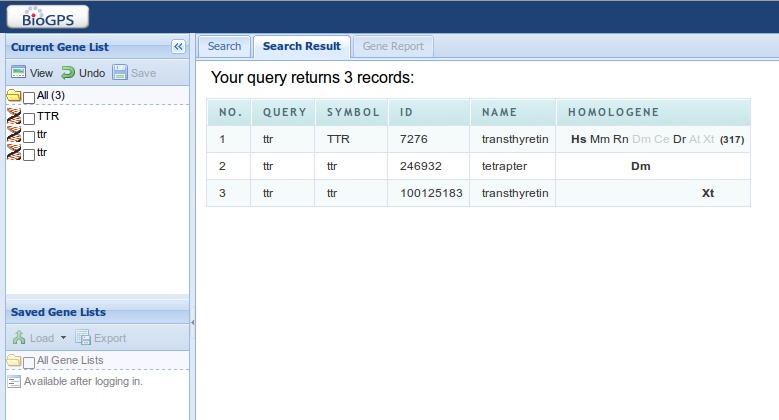
What you should notice: BioGPS finds this gene symbol, and displays the corresponding gene name. It has found three corresponding genes, also listed in the left box as a 'gene list'. We are interested in 'transthyretin', the first. In the column of 'HomoloGene', we see in which species homologues have been found (hover the mouse over the symbols to see more info about the species). Click on the first row to see some info!
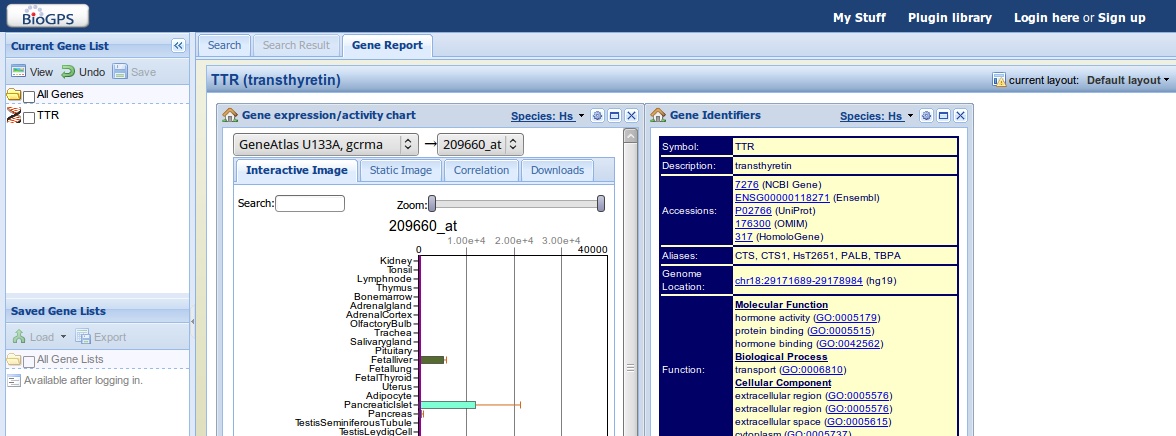
Information is shown in the right window: in this right window, smaller windows display information from different databases. By default, three windows appear, one showing gene expression information, one with gene information and links to relevant databases (of which most of them you should understand - the ones you don't understand get covered in the last presentation :-)), and a final window showing the corresponding wikipedia page. You can drag these windows around, maximise them or delete them.
The power of BioGPS lays in the extensibility of these small windows: different plugins allow you to view more info from other databases. To do so, click on ![]() , which shows you a list of many plugins you might be interested in.
, which shows you a list of many plugins you might be interested in.
Let's show some interaction data: we will display now interaction data from the STRING database. On the BioGPS plugins page, you will find under the 'Most popular' section, the plugin for 'STRING' mentioned.
Click on 'STRING - Proteins and their Interactions': this will take you to a page describing the plugin and the data source. To add it to your current window, click on the blue button 'Add to layout'.

You will now see the STRING info displayed on your page. You can move it around to give it a good place on your overview. We will not go into detail about the data in the STRING database: you can find more on http://string-db.org/ if you are interested.
If you create collection of these views from different databases in BioGPS, you can save these 'views' for later use.
A very interesting feature of BioGPS is that you can quickly browse through homologues in different species. On the top right corner of every windows in BioGPS, there is a dropdown menu with homologues of the current gene. Selecting one of the homologues refreshes the display with information of that homologue. Have a look at the mouse homologue, by selecting it from that little dropdown-menu.
If you want to know more about BioGPS, you can check the help pages on the the BioGPS help pages.
Biomart, get any information you want using Martview
We will show now a very useful tool, called Biomart. We understand now enough concepts to make use of its full potential. Biomart lets you easily retrieve exactly all information that you want. You have GO numbers, and need the corresponding protein sequences of a species? Use Biomart. You have SNP ids from the dbSNP database and want 3'UTR sequences of the genes, use Biomart. It is an indispensable tool to get the date you want.
Biomart - more specifically Martview - is a webbased tool to query databases in a very ingenious way: you set filters = which genes you want to retrieve information for, and you set Attributes = what information you want to have about the genes
Let us start with a simple question: how many proteins in human have the retinol binding domain? Retrieve the list of gene names and descriptions. Instead of going to thé domain database, InterPro, we will use Biomart.
Go to the Biomart homepage. End 2011 the main interface has changed. To access Martview, click on Version 0.7, and then on the link 'Portal'.

Our challenge: we want to retrieve all sequences in human that have the retinol binding domain (IPR002449).
Martview works in three steps:
- select the database you want to query (depends on the type of information you want to retrieve)
- set conditions which you results need to match (can be plenty of them!)
- select what you exactly want to retrieve from the results (e.g. sequence, names, domains, gene ontology terms,...)
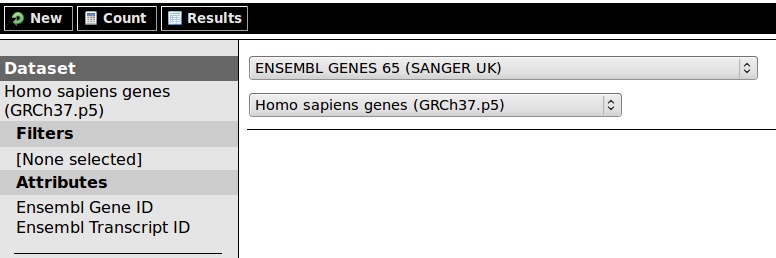
To accomplish this, select Ensembl Genes 65 from the database menu (this database will be covered in the last module).
A new selection appears: restrict here to the 'Homo sapiens genes' dataset.
So, our database we want to search is set: step 1 is finished.
Filters and Attributes appear: both are clickable. For step 2, selecting the conditions which must be met, click on 'Filters'. Under protein domains, fill in IPR002449 (the retinol-binding domain) and set the identifier to InterPro IDs.
When you click on ![]() , you will see how many genes match these conditions, which are
, you will see how many genes match these conditions, which are ![]() .
.
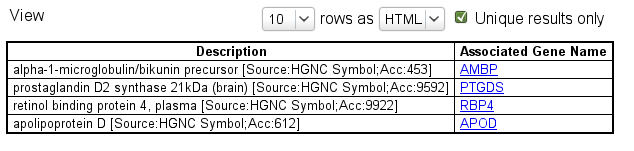
As a last step, which is formatting the output, click on Attributes, open the 'Gene tab' and select 'Description and Associated Gene Name'.
.
Click on ![]() and see how your results are appearing! You may want to select
and see how your results are appearing! You may want to select ![]() .
.
How fast was that? The results from that database are in your hands in the form you want.
| Which functions do the proteins have from the mouse chromosome 12 which are spanning a membrane? |
|---|
| Let's translate: functions, so we can look for Gene Ontology terms, from 'Molecular function'. |
| We need to retrieve those genes: with Biomart should this be no problem! We can fetch for each gene the GO term. |
| So now we need a tool which can somehow summarize the GO information that we fetch from Biomart. |
| To summarize GO terms, you can browse the Gene Ontology tools from www.geneontology.org: on this page you can find the CateGOrizer tool. In short: it counts the occurrence of GO terms and reports statistics of them. |
| So now you have your tools! Can you answer the question? |
Above examples show that Biomart can be very useful to retrieve particular information from a selected set of genes. Biomart can be also a very convenient tool to convert database identifiers of all kind to any other kind.
Convert following affymetrix probe id's (names of probes used on microarrays) to their associated gene names.260649_at 265099_at 255943_at 245070_at 245070_at 258147_at 258147_at 252197_at 246242_at 253188_at |
|---|
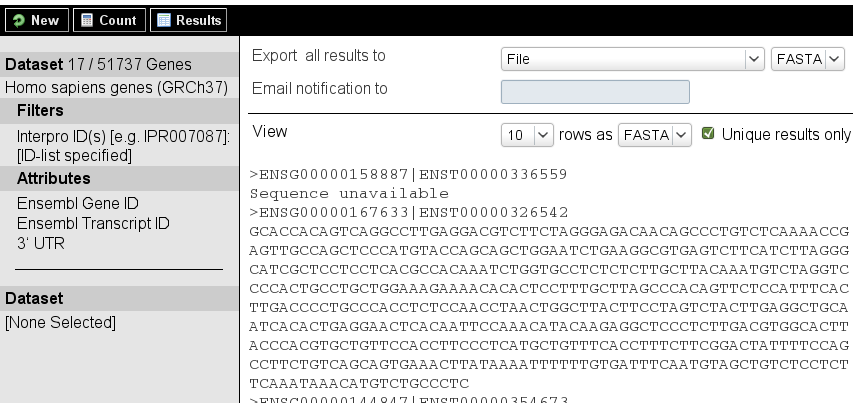
Biomart is also useful for retrieving specific sequences, such as in next task. Can you bring it to a good end?
| Retrieve the 3' UTR sequences of all human genes which have domain IPR020408. |
|---|
| |
Additional exercises
The Gene database at NCBI collect many relevant resources about your gene of interest
| How many types of endothelin receptors are known in Human? |
|---|
| Go to Entrez and enter in the search field: "endothelin receptors AND Human[Organism]" |
| Click through on the Genes database. |
| Another way: go first to the Gene database at NCBI, set limits over there to Homo sapiens and search for Endothelin receptor |
| The official gene name and abbreviation is shown on the records: |
For more information on the HUGO Gene Nomenclature see here.
The PubChem database at NCBI provides information about chemicals and their influence on biological molecules
Entrez contains also a link to databases with chemicals, PubChem. An interesting resource which should be equally user-friendly as other Entrez tables.
Go to parent Basic bioinformatics concepts, databases and tools#Exercises_during_the_training